Research
Educational, Professional & Achievements Details
Structure and dynamics control chemistry. Our broad interest is to gain the physical insight of this control in small molecules (Ultrafast-Spectroscopy) to complex biosystems (Biophysical- Chemistry) and solution (Solution-Chemistry), using established spectroscopic tools and new methodologies (Methodology-Development) as needed.
Ultrafast Spectroscopy
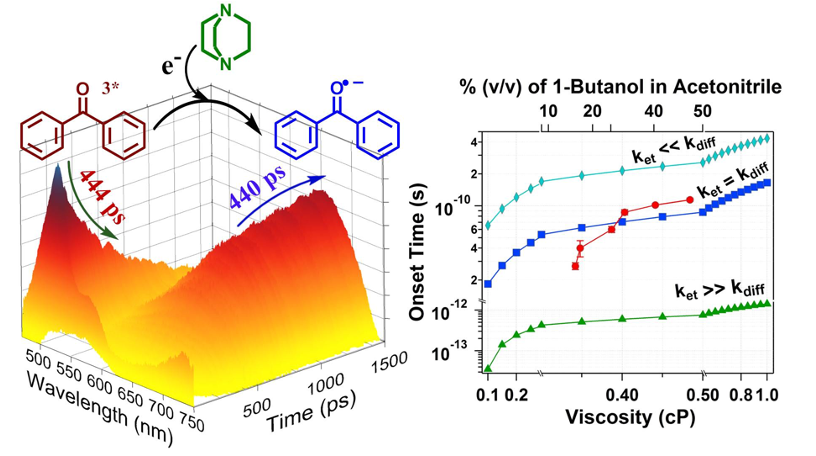
Ultrafast photochemistry and photophysics are our fundamental interest. Traditional photochemistry relies on the fact that internal conversion, vibrational cooling, and solvent relaxation are faster than the radiative transition. The study of photophysics in a timescale faster/comparable to such molecular relaxation provides unique insights. For example, photochemistry in the upper excited state manifold or involving a potential energy barrier might open up new possibilities for wavelength selective chemistry and/or reaction dynamics. We have taken up several related projects. We are basically interested in various photophysical processes e.g., photoinduced large amplitude motion, electron/proton transfer, energy transfer, excimer formation and intersystem crossing. Moreover, to mimic biological conditions, we introduce confinement. Our research can be crudely divided into the following three sub-division
Current research:
- Role of vibration in intersystem crossing process.
- Homo-energy transfer.
- Marcus inversion in excited state proton transfer reaction.
Contact:
Biophysical Chemistry
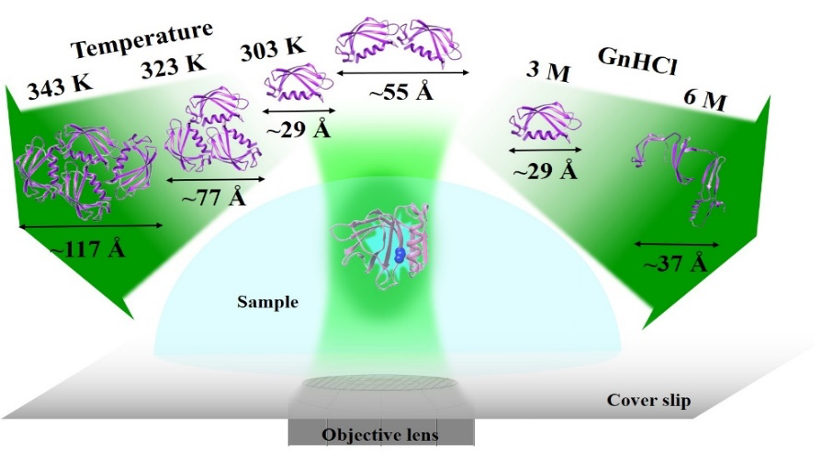
One of our main research goals is to obtain additional insight into the structural, functional and dynamical aspects of proteins. We use various ensemble average (like steady-state and time-resolved emission, CD spectroscopy) and single molecular level spectroscopic techniques (fluorescence correlation spectroscopy) routinely for this. We also use other techniques like AFM, microscopic imaging, HPLC, DLS, etc., as and when required. Another essential part of our research is the site-selective tagging of protein molecules to achieve desired specific information. Crudely, our research interest can be divided into three parts
Current research:
- We are aiming to demonstrate that, contrary to the general belief, the solvent structure (and not the positional restriction due to excluded volume) primarily determine the stability of the protein in the crowded medium
- Role of active-site dynamics in controlling the enzymatic activity
- DES (or more accurately hydrated DES) has already been postulated as the third medium inside a living system that helps to maintain physiological activities even in dehydrated conditions or at sub-freezing temperatures. This is surprising because the presence of water is the crucial factor that controls protein activity. We are aiming to have a physical insight into this
Contact:
Solution Chemistry
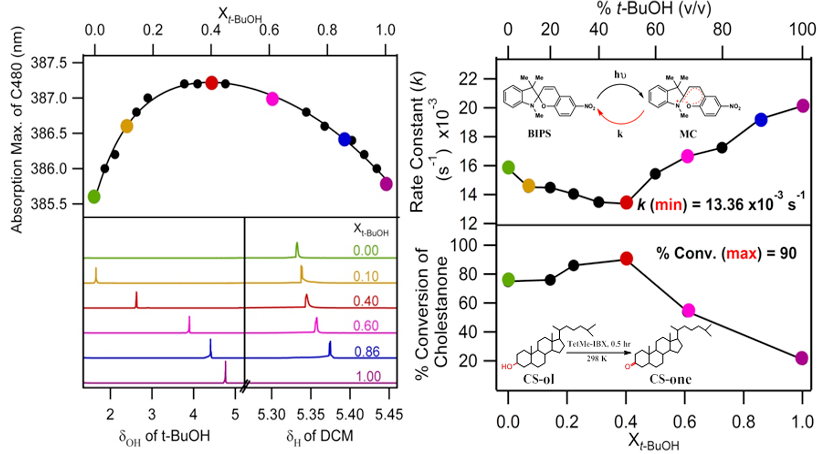
Molecular-level structure and dynamics decide the functionality of a solvent. Therefore, a significant amount of effort was dedicated, being dedicated, and will be dedicated in understanding its structural and dynamical features. Using various steady-state and time-resolved fluorescence measurements and NMR spectroscopy, we try to understand the following
Current research:
- We are aiming for better molecular-level picture of the synergistic solvation in a binary solvent mixture and DES
- Structure and dynamics of the warm-like micelle.
- Understand various forces and interactions responsible for the unique behaviour of binary mixture and DES.
- Synergism in DES and hydrated DES.
Contact:
Materials Chemistry
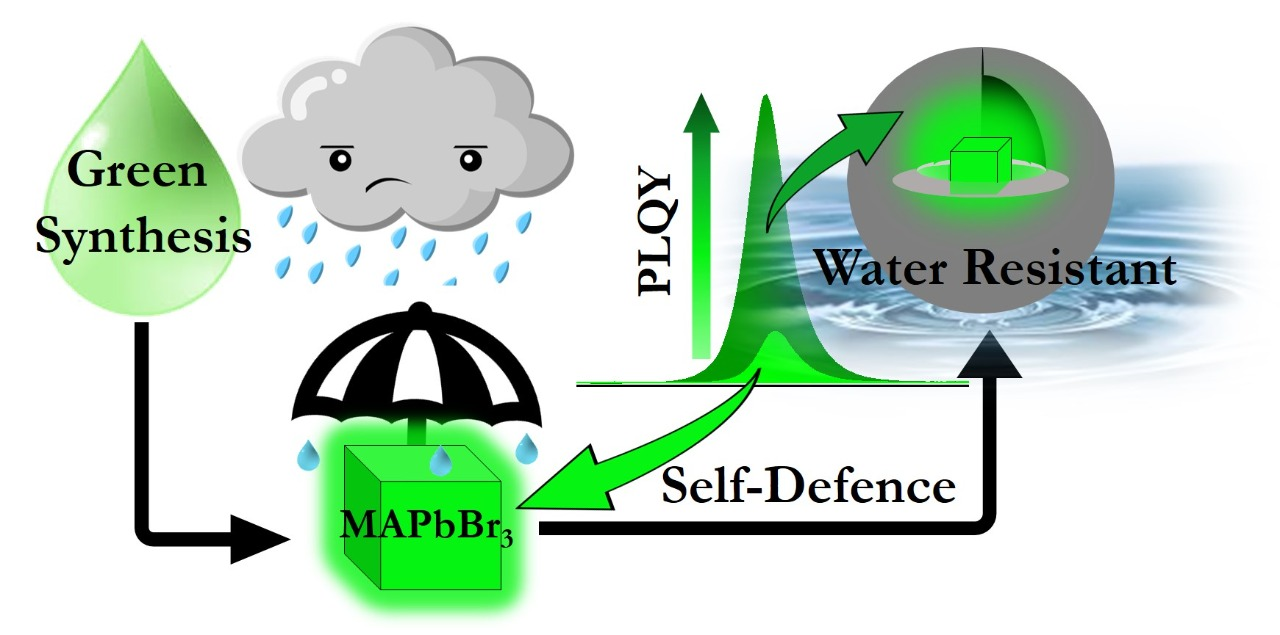
The design and development of new material are indispensable for sustainable growth and is at the heart of chemistry. Though we are not specialists in this area, we try to contribute to this by understanding the material's photophysics and subsequently predicting suitable modifications to have the desired function. We are specifically interested in providing this physical insight.
Current research:
- Inspired by our understanding of the solution chemistry, we are now trying to synthesize photoluminescence, water-stable perovskite nanocrystal highly in a green medium.
- Synthetic and post-synthetic modifications to have better nanocrystals.
- Contemplating the charge carrier dynamics to understand the physical basis of doping.
Contact:
Methodology Development
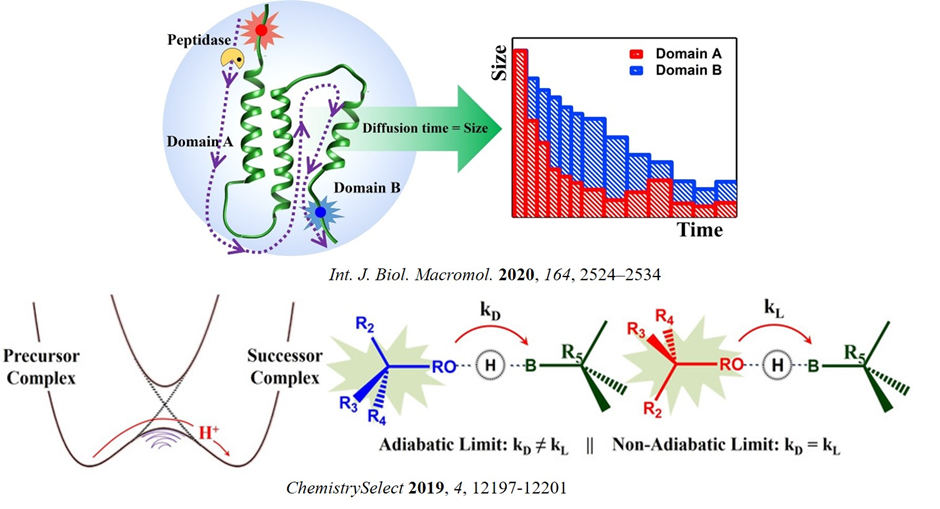
Development of new methodologies and novel approaches necessarily laid the foundation of exciting discoveries in physical chemistry. Recently, we have become interested in this direction. In a recent work, we successfully developed a method to probed the directionality of proteolysis by applying single molecular level fluorescence correlation spectroscopy. Contrary to traditional biophysical methods, minimal sample requirement in this method might provide a significant step forward in elucidating the mechanism of protein digestion (Int. J. Biol. Macromol. 2020, 164, 2524). We also developed a novel way to adjudge the presence of adiabaticity in the proton transfer step of a reaction by exploiting diastereoselectivity. It can overcome the limitations of only available experimental technique, i.e., kinetic isotope effect, to elucidate the proton transfer mechanism. (ChemistrySelect 2019, 4, 12197).
Current research:
- Analytical prediction of viscosity decoupling and dynamic heterogeneity in solvent media.
- Conformational heterogeneity of protein core in its native and denatured state in a domain-specific manner.
Contact:

